Biologia plantarum 70:11-17, 2026 | DOI: 10.32615/bp.2026.002
RT-qPCR analysis of genotype-specific beet yellows virus accumulation in sugar beet
- 1 Plant Virus and Vector Interactions, Czech Agrifood Research Center (CARC), Drnovská 507, 16106 Prague, Czech Republic
- 2 Department of Plant Protection, Faculty of Agrobiology, Food and Natural Resources, Czech University of Life Sciences Prague, Kamýcká 129, 16500 Prague, Czech Republic
Background: Rapid and objective characterisation of viral accumulation requires methods combining fast detection and temporal resolution.
Aims: Develop a two-phase approach for screening sugar beet genotypes for beet yellows virus (BYV) accumulation dynamics.
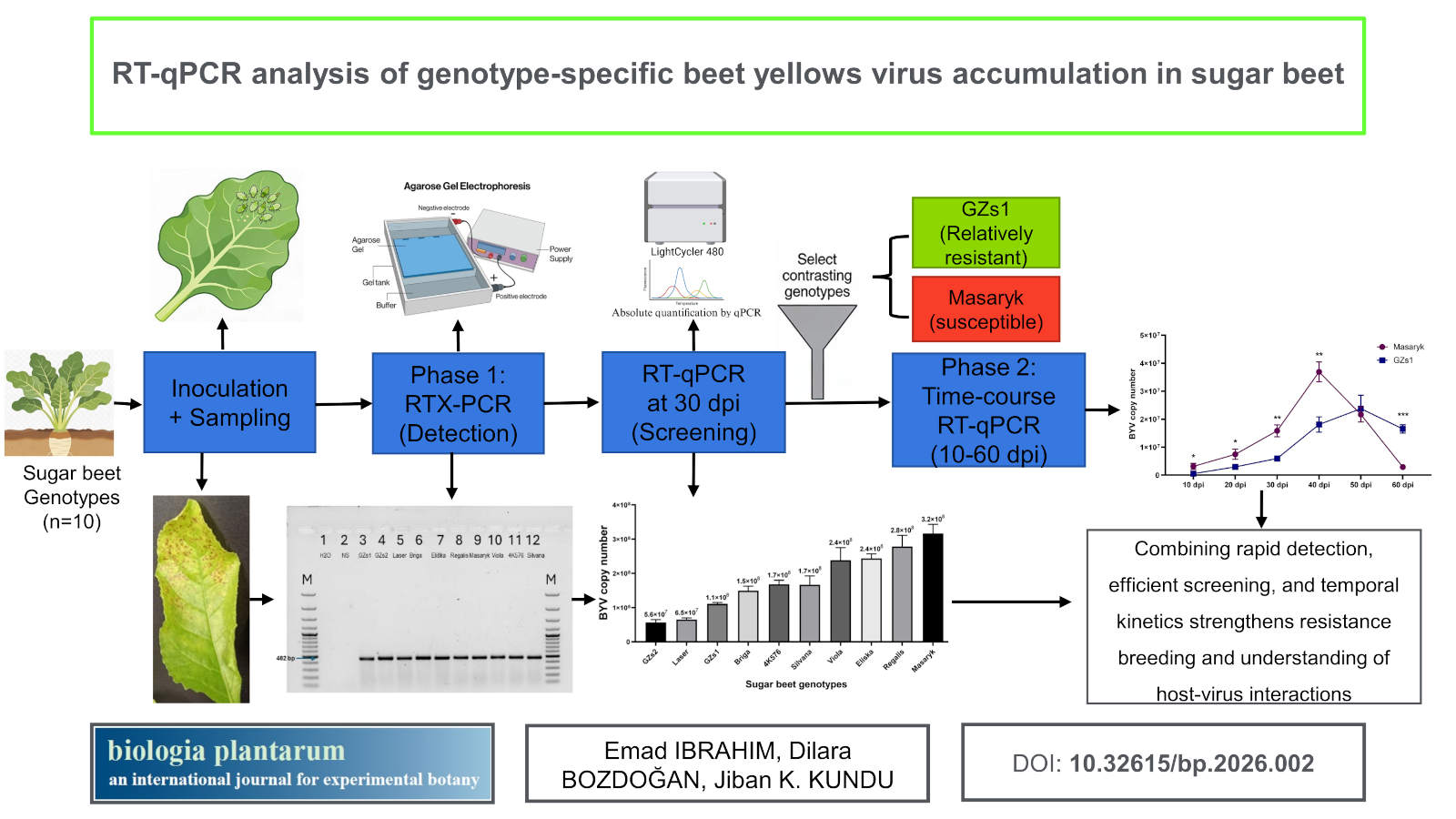
Methods: Phase 1 employed one-enzyme RTX-PCR for rapid BYV detection, followed by single-time-point RT-qPCR at 30 days post-inoculation (dpi) to screen ten genotypes. Phase 2 performed time-course RT-qPCR at six time points (10-60 dpi) on two contrasting genotypes selected from Phase 1, with assay performance validated by a standard curve.
Results: One-enzyme RTX-PCR confirmed BYV infection, enabling quantification; single-time-point RT-qPCR at 30 dpi showed a 5.7-fold titre range (5.6 × 107 to 3.2 × 108 copies), while time-course RT-qPCR revealed distinct trajectories, relatively resistant GZs1 increased gradually (~5.0 × 105 at 10 dpi to 1.7 - 2.3 × 107 at 40 - 60 dpi) whereas susceptible Masaryk rose rapidly early (~3.1 × 106 at 10 dpi) and peaked at 3.7 × 107 by 40 dpi, dynamics not captured by single-time-point measurements; standard curve metrics indicated high assay quality (R2 = 0.9976; efficiency = 103%).
Conclusions: This two-phase method combines speed with precision for effective genotype comparison. It reveals BYV accumulation dynamics that are not captured by single‑point assays.
Keywords: Beet yellows virus, kinetics, RT-qPCR, RTX-PCR, sugar beet, viral quantification.
Received: December 22, 2025; Revised: February 2, 2026; Accepted: February 4, 2026; Published online: February 23, 2026 Show citation
Supplementary files
| Download file | 7318_Ibrahim_Suppl.pdf File size: 169.69 kB |
References
- Alquicer, G., Ibrahim, E., Maruthi, M.N. & Kundu, J.K. (2023) Identifying putative resistance genes for barley yellow dwarf virus-PAV in wheat and barley. Viruses, 15, 716.
 Go to original source...
Go to original source... - Bass, C., Puinean, A.M., Zimmer, C.T. et al. (2014) The evolution of insecticide resistance in the peach potato aphid, Myzus persicae. Insect Biochemistry and Molecular Biology, 51, 41-51.
 Go to original source...
Go to original source... - Baulcombe, D. (2004) RNA silencing in plants. Nature, 431, 356-363.
 Go to original source...
Go to original source... - Bertinelli, G., Tizzani, L., Luigi, M., Monticelli, S. & Ilardi, V. (2024) Development and validation of one-step reverse transcription-droplet digital PCR for plum pox virus detection and quantification from plant purified RNA and crude extract. Plants, 13, 3276.
 Go to original source...
Go to original source... - Borgolte, S., Varrelmann, M. & Hossain, R. (2024) Time point of virus yellows infection is crucial for yield losses in sugar beet, and co-infection with beet mosaic virus is negligible under field conditions. Plant Pathology, 73, 2056-2070.
 Go to original source...
Go to original source... - Carroll, J.E., Smith, D.M. & Gray, S.M. (2016) Preferential acquisition and inoculation of PVYNTN over PVYO in potato by the green peach aphid Myzus persicae (Sulzer). Journal of General Virology, 97, 797-802.
 Go to original source...
Go to original source... - Cheng, X. & Wang, A. (2017) Multifaceted defense and counter-defense in co-evolutionary arms race between plants and viruses. Communicative & Integrative Biology, 10, e1341025.
 Go to original source...
Go to original source... - Ding, S.-W. (2023) Transgene silencing, RNA interference, and the antiviral defense mechanism directed by small interfering RNAs. Phytopathology, 113, 616-625.
 Go to original source...
Go to original source... - Dolja, V.V. (2003) Beet yellows virus: the importance of being different. Molecular Plant Pathology, 4, 91-98.
 Go to original source...
Go to original source... - Dráb, T., Svobodová, E., Ripl, J. et al. (2014) SYBR Green I based RT-qPCR assays for the detection of RNA viruses of cereals and grasses. Crop and Pasture Science, 65, 1323-1328.
 Go to original source...
Go to original source... - Ellefson, J.W., Gollihar, J., Shroff, R., Shivram, H., Iyer, V.R. & Ellington, A.D. (2016) Synthetic evolutionary origin of a proofreading reverse transcriptase. Science, 352, 1590-1593.
 Go to original source...
Go to original source... - Garcia-Ruiz, H. (2018) Susceptibility genes to plant viruses. Viruses, 10, 484.
 Go to original source...
Go to original source... - Hoffmeisterová, H., Kratochvílová, K., Čeřovská, N. et al. (2022) One-enzyme RTX-PCR for the detection of RNA viruses from multiple virus genera and crop plants. Viruses, 14, 298.
 Go to original source...
Go to original source... - Hossain, R., Menzel, W., Lachmann, C. & Varrelmann, M. (2021) New insights into virus yellows distribution in Europe and effects of beet yellows virus, beet mild yellowing virus, and beet chlorosis virus on sugar beet yield following field inoculation. Plant Pathology, 70, 584-593.
 Go to original source...
Go to original source... - Hossain, R., Willems, G., Wynant, N., Borgolte, S., Govaerts, K. & Varrelmann, M. (2024) Aphid-mediated beet yellows virus transmission initiates proviral gene deregulation in sugar beet at early stages of infection. PLoS ONE, 19, e0311368.
 Go to original source...
Go to original source... - Huang, J., Chen, Y., Zhu, X. et al. (2025) Viral detection in Phalaenopsis orchids using high-throughput sequencing and one-step multiplex RT-PCR. Plant Disease, 109, 2257-2261.
 Go to original source...
Go to original source... - Ibrahim, E., Rychlá, A., Alquicer, G. et al. (2023) Evaluation of resistance of oilseed rape genotypes to turnip yellows virus. Plants, 12, 2501.
 Go to original source...
Go to original source... - Jebakumar, R.M., Balasubramanian, V. & Selvarajan, R. (2018) Virus titre determines the efficiency of Pentalonia nigronervosa (Aphididae: Hemiptera) to transmit banana bunchy top virus. VirusDisease, 29, 499-505.
 Go to original source...
Go to original source... - Jiménez, J., Tjallingii, W.F., Moreno, A. & Fereres, A. (2018) Newly distinguished cell punctures associated with transmission of the semipersistent phloem-limited beet yellows virus. Journal of Virology, 92, e01076-18.
 Go to original source...
Go to original source... - Kanapiya, A., Amanbayeva, U., Tulegenova, Z. et al. (2024) Recent advances and challenges in plant viral diagnostics. Frontiers in Plant Science, 15, 1451790.
 Go to original source...
Go to original source... - Kang, B.-C., Yeam, I. & Jahn, M.M. (2005) Genetics of plant virus resistance. Annual Review of Phytopathology, 43, 581-621.
 Go to original source...
Go to original source... - Kumar, R., Kaundal, P., Tiwari, R.K. et al. (2022) Establishment of a one-step reverse transcription recombinase polymerase amplification assay for the detection of potato virus S. Journal of Virological Methods, 307, 114568.
 Go to original source...
Go to original source... - Kumar, R., Kumar Lal, M., Prasad, P. & Tiwari, R.K. (2023) Editorial: Current advancements in real-time plant pathogen diagnostics: from lab assays to in-field detection. Frontiers in Plant Science, 14, 1255654.
 Go to original source...
Go to original source... - Lee, C., Kim, J., Shin, G.S. & Hwang, S. (2006) Absolute and relative QPCR quantification of plasmid copy number in Escherichia coli. Journal of Biotechnology, 123, 273-280.
 Go to original source...
Go to original source... - Lough, G., Kyriazakis, I., Bergmann, S., Lengeling, A. & Doeschl-Wilson, A.B. (2015) Health trajectories reveal the dynamic contributions of host genetic resistance and tolerance to infection outcome. Proceedings of the Royal Society B: Biological Sciences, 282, 20152151.
 Go to original source...
Go to original source... - Ma, X., Zhou, Y., Wu, L. & Moffett, P. (2024) Resistance gene Ty-1 restricts TYLCV infection in tomato by increasing RNA silencing. Virology Journal, 21, 256.
 Go to original source...
Go to original source... - Nancarrow, N., Aftab, M., Hollaway, G., Rodoni, B. & Trębicki, P. (2022) Symptomless turnip yellows virus infection causes grain yield loss in lentil and field pea: A three-year field study in south-eastern Australia. Frontiers in Plant Science, 13, 1049905.
 Go to original source...
Go to original source... - Nandudu, L., Sheat, S., Winter, S., Ogbonna, A., Kawuki, R. & Jannink, J.-L. (2024) Genetic complexity of cassava brown streak disease: insights from qPCR-based viral titre analysis and genome-wide association studies. Frontiers in Plant Science, 15, 1365132.
 Go to original source...
Go to original source... - Nithya, K., Parameswari, B., Kumar, S. et al. (2023) Prospecting true ScYLV resistance in Saccharum hybrid parental population in India by symptom phenotyping and viral titre quantification. 3 Biotech, 13, 125.
 Go to original source...
Go to original source... - Pagán, I. & García-Arenal, F. (2018) Tolerance to plant pathogens: theory and experimental evidence. International Journal of Molecular Sciences, 19, 810.
 Go to original source...
Go to original source... - Paudel, D.B. & Sanfaçon, H. (2018) Exploring the diversity of mechanisms associated with plant tolerance to virus infection. Frontiers in Plant Science, 9, 1575.
 Go to original source...
Go to original source... - Pérez-Cañamás, M., Taliansky, M. & Hernández, C. (2022) A viral suppressor of RNA silencing may be targeting a plant defence pathway involving fibrillarin. Plants, 11, 1903.
 Go to original source...
Go to original source... - Pfaffl, M.W. (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Research, 29, e45.
 Go to original source...
Go to original source... - Quenouille, J., Montarry, J., Palloix, A. & Moury, B. (2013) Farther, slower, stronger: how the plant genetic background protects a major resistance gene from breakdown. Molecular Plant Pathology, 14, 109-118.
 Go to original source...
Go to original source... - Ranabhat, N.B., Bruce, M.A., Fellers, J.P. & Rupp, J.L.S. (2022) A reproducible methodology for absolute viral quantification and viability determination in mechanical inoculations of wheat streak mosaic virus. Tropical Plant Pathology, 47, 553-561.
 Go to original source...
Go to original source... - Sharaf, A., Nuc, P., Ripl, J. et al. (2023) Transcriptome dynamics in Triticum aestivum genotypes associated with resistance against the wheat dwarf virus. Viruses, 15, 689.
 Go to original source...
Go to original source... - Smith, H.G. & Hallsworth, P.B. (1990) The effects of yellowing viruses on yield of sugar beet in field trials, 1985 and 1987. Annals of Applied Biology, 116, 503-511.
 Go to original source...
Go to original source... - Soosaar, J.L.M., Burch-Smith, T.M. & Dinesh-Kumar, S.P. (2005) Mechanisms of plant resistance to viruses. Nature Reviews Microbiology, 3, 789-798.
 Go to original source...
Go to original source... - Stevens, M., Hallsworth, P.B. & Smith, H.G. (2004) The effects of Beet mild yellowing virus and Beet chlorosis virus on the yield of UK field-grown sugar beet in 1997, 1999, and 2000. Annals of Applied Biology, 144, 113-119.
 Go to original source...
Go to original source... - Sun, B., Li, S., Pi, Z., Wu, Z. & Wang, R. (2024) Assessment of genetic diversity and population structure of exotic sugar beet (Beta vulgaris L.) varieties using three molecular markers. Plants, 13, 2954.
 Go to original source...
Go to original source... - Svec, D., Tichopad, A., Novosadova, V., Pfaffl, M.W. & Kubista, M. (2015) How good is a PCR efficiency estimate: Recommendations for precise and robust qPCR efficiency assessments. Biomolecular Detection and Quantification, 3, 9-16.
 Go to original source...
Go to original source... - Vegvari, C., Hadjichrysanthou, C., Cauët, E. et al. (2016) How can viral dynamics models inform endpoint measures in clinical trials of therapies for acute viral infections? PLoS ONE, 11, e0158237.
 Go to original source...
Go to original source... - Withycombe, J., Han, J., MacWilliams, J., Dorn, K.M., Nalam, V.J. & Nachappa, P. (2024) Transcriptomic profiling reveals distinct responses to beet curly top virus (BCTV) infection in resistant and susceptible sugar beet genotypes. BMC Genomics, 25, 1237.
 Go to original source...
Go to original source... - Zvereva, A.S. & Pooggin, M.M. (2012) Silencing and innate immunity in plant defense against viral and non-viral pathogens. Viruses, 4, 2578-2597.
 Go to original source...
Go to original source...



